Method
RNA-Seq was used to detect and enumerate DEGs following treatment with both modified and unmodified siRNA, assessing the extent of mRNA upregulation or downregulation in off-target genes.
Results
- Table 1: Modified siRNA (siRNA93-GS) significantly reduced the number of DEGs compared with unmodified siRNA.
- Figures 1A/B: Unmodified siRNA caused widespread changes in gene transcription, affecting multiple genes.
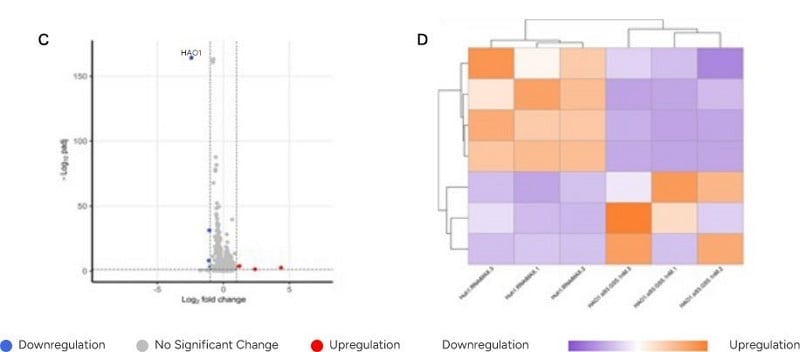
- Figures 1C/D: Modified siRNA (siRNA93-GS) greatly reduced these transcriptional changes.
- Figures 1A/C: Data point HAO1 showed similar mRNA changes along the horizontal axis, indicating that modified siRNA maintained target knockdown efficiency comparable to unmodified siRNA.
Table 1: Modified siRNA (siRNA93-GS) significantly reduced the number of DEGs compared with unmodified siRNA.
| Target Gene | Sample | RNA-Seq Detection | Type I: Homologous off-target genes prediction | Type II: miRNA-like similar off-target genes prediction | ||||
|---|---|---|---|---|---|---|---|---|
| DEGs | AS_blast | SS_blast | AS_mirna_off-target | SS_mirna_off-target | Total mirna_off-target | Down regulated off-target | ||
| HAO1 | siRNA93 | 58 | 0 | 0 | 26 | 6 | 28 | 23 |
| siRNA93-GS | 7 | 0 | 0 | 1 | 0 | 1 | 1 | |
Off-Target Effects of Unmodified siRNA - Gene Transcription Changes
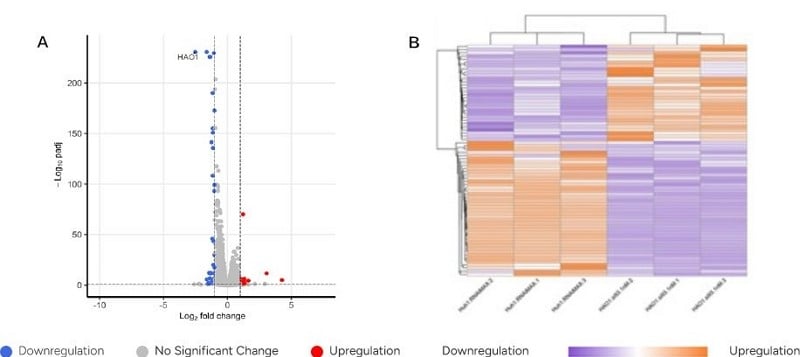
Off-Target Effects of GenScript’s Proprietary Modified siRNA – Gene Transcription Changes